Undergraduate Research Projects
Below are undergraduate research projects completed recently by members of the Rinker-Ross School of Health Sciences. Click images for full-size versions of research posters.
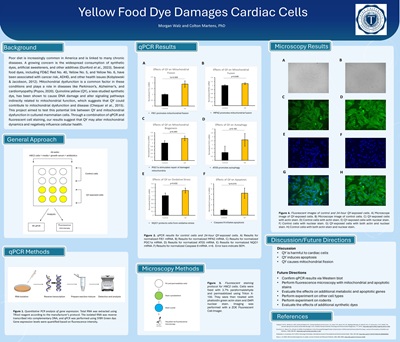
Yellow Food Dye Damages Cardiac Cells, Morgan Walz, supervised by Colton Martens, Ph.D.
Morgan was interested in determining if quinoline yellow—a common dye used in foods and cosmetics—damages cardiac cells. She exposed rat cardiomyocytes to quinoline yellow in a petri dish for 24 hours and assessed its effects on cellular health by measuring gene expression. She found that quinoline yellow damaged the mitochondria and promoted cell death, suggesting that this food dye might have underappreciated health consequences.
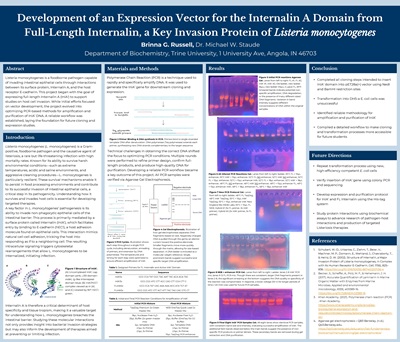
Development of an Expression Vector for the Internalin A Domain from Full-Length Internalin, a Key Invasion Protein of Listeria monocytogenes, Brinna Russell, supervised by Mike W. Staude, Ph.D.
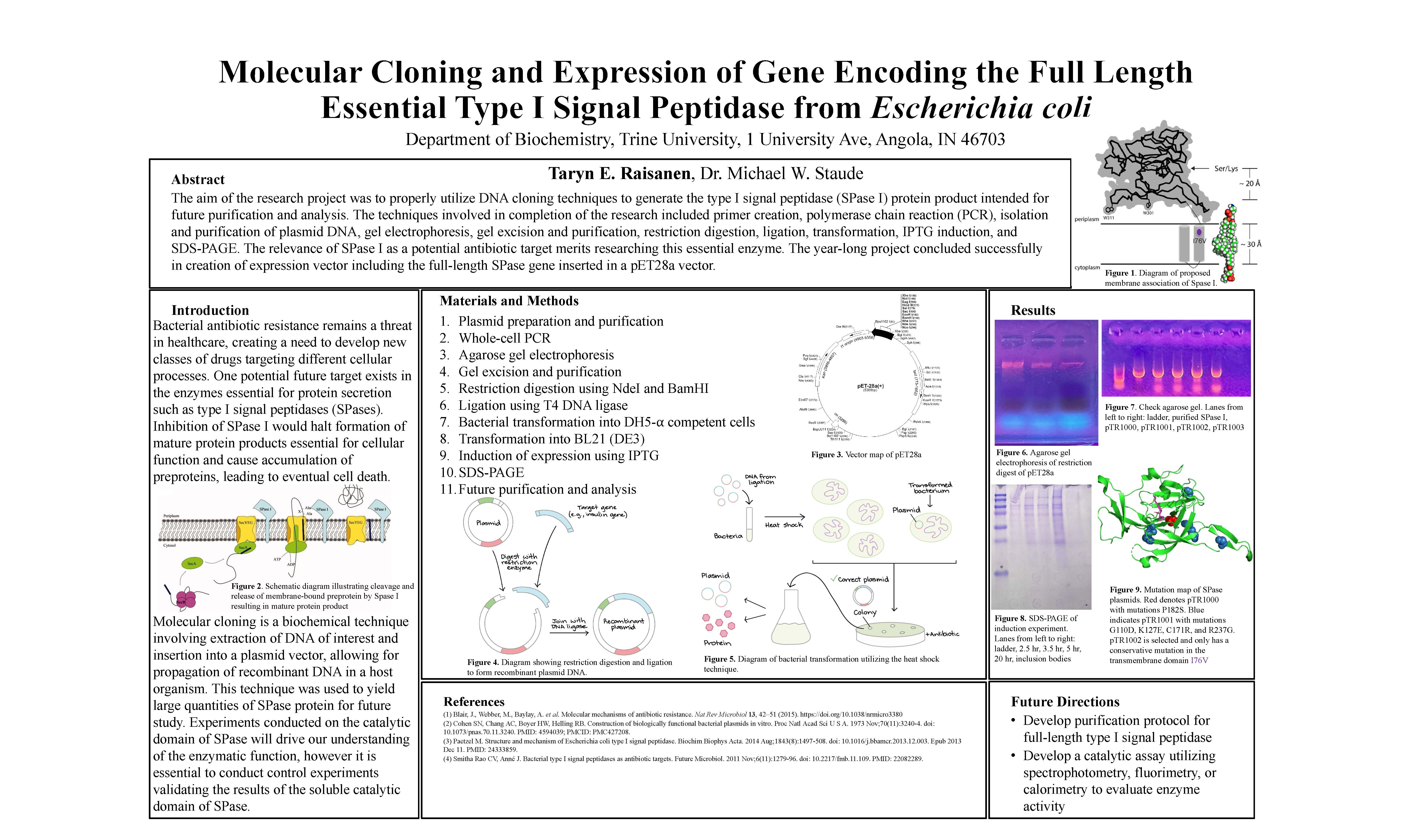
Molecular Cloning and Expression of Gene Encoding the Full Length Essential Type I Signal Peptidase from Escherichia coli, Taryn Raisanen, supervised by Mike W. Staude, Ph.D.
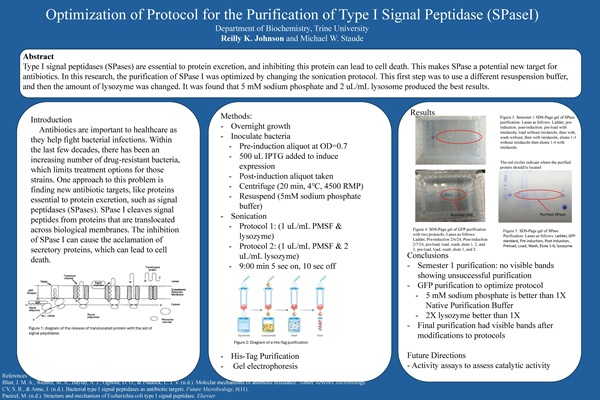
Optimization of Protocol for the Purification of Type I Signal Peptidase (SPaseI), Reilly Johnson, supervised by Mike W. Staude, Ph.D.